Pillars for a global bioeconomy #1: decode
History, markets and future of genomic sequencing in LatAm and the world

For thousands of years, questions like “Who am I? where do I come from? Where am I going?" or “who do I want to be with?” had remained philosophical questions, mostly irrelevant to science, without an objective answer. Later, the industries of the 20th century, based mainly on synthetic chemistry1, created today’s world’s biggest problems, some of which we know as obesity and the ecological crisis.
As any inhabitant of the 21st century would be interested in knowing, the intersection of biological and computational technologies is providing answers to these questions and sustainable solutions to these problems. Not only have we succeeded in sequencing entire human genomes, we've not only reinvented the way we grow meat: we've made these technologies >10x more accesible, in less than 10 years2 🦄.
A few days ago I was talking with the guys from GRIDX—the largest biotech company builder in Latam—and we agreed that there are many more startups of this kind whose story deserves to be told.
In this series of articles, my mission is to give you a holistic overview of the most important startups in the industry, as well as the 4 technologies that will be pillars in the creation and development of the world’s new bioeconomy, starting with sequencing.
The largest hackathon in human history
32 years ago, one of the most ambitious scientific projects in history to date began: decoding the human genome. The Human Genome Project (HGP) is famous for having had a total cost of 3 billion dollars and a duration of thirteen years.
Both collaborative and competitive, with funding from the NIH and the US Department of Energy. A consortium was formed with researchers at 20 universities in the UK, Japan, France, Germany and China. Seven years before completion, the Celera company (founded by Craig Venter) announced that it could complete the project in as little as 3 years and at a much lower cost. Wondering how?
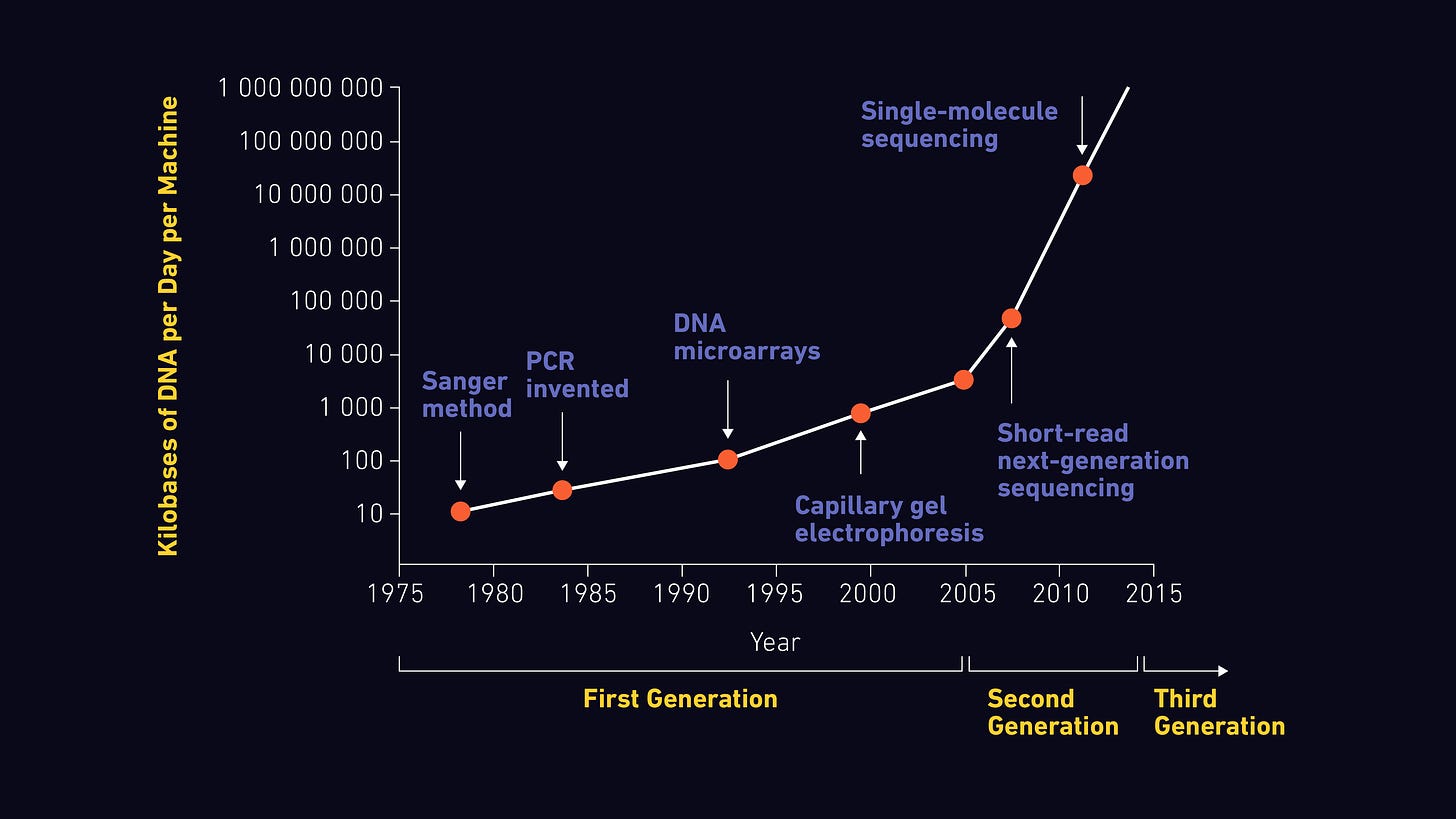
Before having today’s sequencing technologies, we went through many methods, “Sanger” being the most popular one. In a nutshell, Sanger seq consists of 3 main steps: separating the DNA double helix, replicating it using fluorescently labeled bases, using capillary gel electrophoresis to separate the copies according to their length, from shortest to longest. As they pass through the gel, the color of their terminal base is detected, thus obtaining a continuous sequence.
Interestingly, Celera’s real advantage, had not to do with sequencing per se, but rather with “pre-processing”. Although both groups used shotgun sequencing, (fragmenting DNA into smaller pieces), the big difference is that the type of shotgun seq used by the HGP required cloning pieces of DNA into bacteria and then sequencing it, which meant more financial and time resources, in contrast to the Celera method which could sequence directly3.

Although this race ended in a tie, we ought to acknowledge that neither of the teams sequenced an actually complete genome then (only the 90-92% which is not strongly bound by proteins). It was not until 2022 that the entire sequence was completed once and for all, including the famous ends of the chromosomes, the telomeres.
A new sequencing era
Although revolutionary and still widely used (90% of the world's sequence data is produced this way), we know that the Sanger method is not the most efficient when we want to sequence complete genomes at the most affordable price. That is why, in recent decades, other methods have emerged that fall into the category of Next Generation Sequencing (NGS).
Roughly speaking, NGS can sequence millions of pieces of DNA at a time, so it is faster for larger sequences, it can give us more data with the same amount of DNA, and it is better at detecting variants.
Here are 5 NGS technologies that are changing the game…
1/ SBS > SBF: the most popular method? 🤔
Sequencing by synthesis (SBS) is the most widely used method, including by companies such as Illumina, GenapSys, and Thermofisher. The recipe for sequencing DNA through this method goes like this:
Prepare the library: fragment the DNA to be sequenced and add adapters to amplify it via bridge amplification. Add “tags” to assemble it later.
Sequence: add nucleotides marked with fluorescent proteins to be able to detect the sequence of each piece of DNA.
Analyze: time to put the fragments together through bioinformatics!
2/ Nanoballs > Pokeballs: US vs China for the $100 genome 🥊
The size of the sequencing market is over $4 billion dollars and growing at a CAGR of 11.4%4. Though multiple reports indicate that Illumina is the majority player with up to 80% market share5, we must remember that we are talking about the young kingdom of biotechnology, where the crown and throne belong to that who conceives the most magical technology.
More than 30 years ago, the Chinese group BGI (Beijing Genomics Institute) was one of the collaborators in the human genome project. In a short time, they have become both Illumina's largest customer and fiercest competitor. This is what Jay Flatley (former CEO of Illumina) had to say in 2013 regarding BGI's acquisition of Complete Genomics:
Yes, they are a very significant customer of ours. We want to maintain a great relationship with them. But we’re not sure it’s in the U.S. national interest to sell the formula for Coke. It’s different when people just buy Coke6.
Complete Genomics was founded by Serbian scientist Radoje Drmanac, who is said to have been one of the brightest developers of new genomic technologies, with more than 4 co-founded companies. Because Complete needed more investment and couldn't compete with Illumina yet, they accepted BGI's takeover offer for $123 million—surely a bargain for what they'll end up achieving.
What Illumina feared, what makes the DNBSEQ method superior, is replacing the use of PCR with a more precise amplification method called Rolling Circle Amplification. Because the original DNA fragment is always used to make copies, errors are almost impossible and significant savings in reagent cost are achieved.
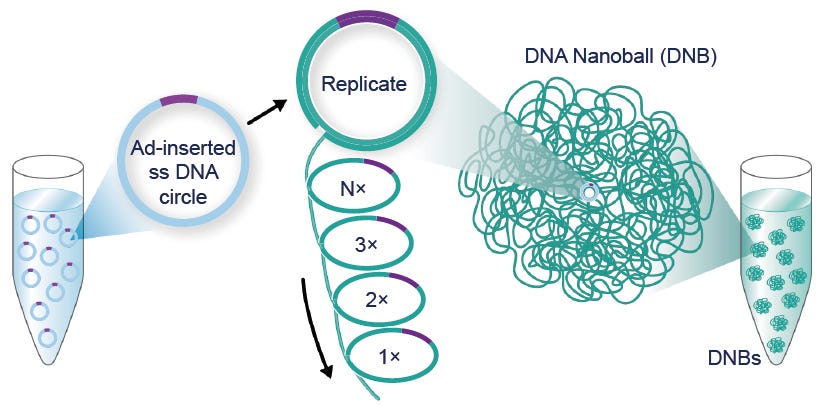
After more R&D, BGI announced in 2020 that it could sequence a whole human genome for just $1007. It wasn't long before Nebula Genomics, an American direct-to-consumer genetic testing company, offered to sequence whole genomes for just $300 USD.
And just a few months ago, Illumina launched its counterattack with the beautifully and futuristic-looking Illumina X Sequencer, which can do whole genome seq for $2008.

3/ Sequencing CD > Normal CD: engineering to 10x sequencing 💿
Also in 2022, Ultima Genomics announced that it could also sequence whole genomes for $100 USD9. Realizing that the majority of sequencing costs come from the flow cell ("chip") and reagents, Ultima developers created a new circular design that spins like a CD. In addition to not using as many fluorescent bases, they use neural networks to assemble the sequence afterwards. They sequence cheaper ($1/Gb) and faster10.

4/ Pyrosequencing > pyrotechnics: those biotech tricks 🎆
For this recipe we will use two special ingredients: APS and luciferin. Every time a complementary nucleotide is added to our sequence of interest, pyrophosphates (Ppis) will be released, which will use the APS to convert into ATP. This will convert luciferin to oxyluciferin, which emits light. This light is detected every base and thus DNA is sequenced.

Something similar happens in the Ion Semiconductor Seq technology, but instead of detecting pyrophosphates, the pH is measured as hydrogen ions are released.
5/ MinION > Minion: the iPhone of sequencers 📱
And for hobbyists like me, there's the iconic minION: the first pocket DNA sequencer that costs the same as an iPhone ($1,000 USD) and plugs right into your laptop 😍.
Oxford Nanopore, the company behind the minION, is named after the place it was founded and the Nanopore technology it has developed. Instead of using fluorescent bases, they pass the DNA through 2-nm-diameter pores, which identify the current that each base carries. With this 4th generation technique, or Single Molecule Sequencing (SMS), readings of 1 million bases can be made in real time and without amplification.
Though Nanopore outperforms all the others in per hour output, that PacBio (4th generation seq) has proven to have greater continuity, correctness and completeness scores, at a lower price than both Nanopore and Illumina. Maybe it’s them who we should be looking at! 👀

Decoding LatAm
The aforementioned advances in NGS sequencing technologies—as well as many others including lab-on-a-chip technology and electron microscopy1112—can only mean one thing: biotech builders don't follow Moore's Law… but exceed it!

Image A shows us how new technologies are being adopted increasingly faster. Today, more than 50% of the world's population has access to magical technology (smartphones and the internet) to do things like video calls thousands of miles away or learn literally anything. My big question is: how long will it be before we can have outstanding healthcare tools at our fingertips?
These are 5 startups that apply sequencing technologies to make that vision a reality. 4 of them were proudly started in Latam, 3 are part of GRIDX’s portfolio and one sneaked in as a Bostonian company.
1/ ArgenTAG
We are building the first truly scalable single-cell solution for long read sequencers. To understand the complexity of human biology, and improve people's lives—Leandro Ciappina, COO.
What if we could make real-time, single-cell, long-read sequencing more accurate AND accessible? Sequencing errors increase as the number of cells to be sequenced increases. Although companies like 10x genomics and Parse Biosciences also work with single-cell, they only do short-read sequencing.
ArgenTAG has created 2 products to solve this problem: a “barcode” kit and software for post-sequencing processing. The investigator:
Receives the ArgenTAG nucleotide tagging kit (NS-watermark)
Follows the protocol created by ArgenTAG to tag genetic material without needing special equipment.
Reads the results in ArgenTAG's proprietary software.
This team of professionals, together with George Church as their scientific advisor, offers its clients longer and more accurate readings, at up to 20x less cost. During their acceleration in the renowned IndieBio program, they managed to mix 100k cells, label and sequence them, and recover the complete profile.
2/ Bitgenia
As a leader in pharmacogenomics in LatinAm, Bitgenia takes clinical analysis to the genetic level. Sequence entire exomes and genomes, analyzing more than 20 thousand genes related to hereditary cancer, cardiology, reproductive genetics, neurology and practically any other medical specialty.
The process is simple:
Health professionals request the test from their patients and work together with Bitgenia to evaluate the study to be carried out.
The patient sends a saliva sample to Bitgenia, who sequences and analyzes it on their bioinformatics platform, B_Platform.
Bitgenia gives the doctor a report with the analysis and classification of variants associated with the patient's symptoms.
Did you know that most drugs are effective on only 25-75% of the patients to whom they are administered? Bitgenia has collaborated with leading pharmacology companies such as Novartis for the diagnosis of autoinflammatory diseases, GSK to generate a personalized treatment against ovarian cancer, and AstraZeneca for oncology diagnostic projects.
3/ Cryosmetics
Well, not only have a gut microbiome. Many years ago, our skin formed a symbiosis with the microbes in the environment; while we give them certain nutrients to survive, they protect us from pathogens and interact with our immune system, helping to keep our skin healthy.
Unfortunately and ironically, the highly processed products used today actually hurt our skin microbiome. That is why Cryosmetics' mission is to improve your skin health through a skin microbiome (16s RNA seq) screening and a personalized treatment using 100% natural products.
Fair, nowadays anyone can use “100% natural” on their labels. You just need to know that among the ingredients that Enteria uses are the highest quality vegetable oils, such as avocado, tomato seed and cranberry oil.
Ready to meet your multitudes?
4/ Nebula Genomics
23 & *who? Ancestry who?—If I made a list of all the things I wished for in a genetic testing service, the result would be Nebula. Not only do they sequence your entire genome at the lowest price to the consumer, they don't just use that information to tell you about your ancestry, personality, possible diseases, and ideal diet and physical activity…
*sorry, I just wanted to sound cool. You guys rock 23&Me and Ancestry! 🐐
One of its biggest value propositions here is to give you control over this information through blockchain technology. In theory, this means that you have complete power over who has access to your genetic information. It is up to you if you want to keep it private and only for yourself, or if you want to share it with researchers in which case, your "genetic profile" would be updated according to new discoveries.
At one point, they even had the option of sequencing your genome completely for free, in exchange for "donating" that information for research—we couldn't expect less from one of the more than 20 biotech startups co-founded by renowned researcher George Church.
5/ Omica
Building a biobank to give latinos power over their genetic data, aligning incentives with local institutions to develop precision medicines through that data. Podcast episode (in Spanish) with the founder here. More on Omica soon…
AUG (just the beginning)
Let’s analyze the 2 breeds of biostartups we’ve seen so far:
B2C: attractive products, replicate proven models, less risky, founders from almost any background, penetrating new market with existing tech.
B2L (business to lab) or B2B: unsexy products, mostly novel, riskier, experienced researchers spinning out from universities, developing products for an existing market (take ArgenTAG who are building in the US now).
Though I idolater the idea of building something brand new, I also see a huge opportunity for healthcare B2Cs here. Just as Rappi and Beek have replicated UBER and Audible, we can create more options for bioproducts here.
I emphasize healthcare because I have a loosely held hypothesis: LatAm’s unfair advantage to create magical technology actually lies in our megabiodiversity. Not only do we have some of the most diverse human genomes worldwide. According to UNEP, 60% of the world’s terrestrial species can be found here. Still, less than 0.5% of plant species in the world have been fully sequenced—more on this soon! ; )
Understanding the importance of this diversity, Illumina has opened 3 genomic centers in Colombia, Brazil and Mexico13. They’re making Illumina equipment available to researchers to use here while getting access to data of even hundreds of thousands of latinx genomes.
As we immerse into the century of biology, we will have to think of who, if anyone, owns biological data14 and make sure it's used for the benefit of humankind. After all, the new global bioeconomy is not only about well, business and biotech. It’s also about assuming a moral responsibility for the biodiversity that we have in each region, country and community; to understand that each cell guards the answers to the world’s biggest problems, to the most ancestral questions; it’s a journey to the past, present and future of our planet, Earth.
May we all use our biological heritage to grow a 10x better world. A new global bioeconomy is emerging and we are all helping to grow it. Wanna join us? 🧬 🌎
Full genome sequencing was $1,000 USD in 2015, China’s BGI announced que it could do that for $100 in 2020. Producing the first lab-grown burger was $325,000 USD in 2013, today the price is less than $8 USD for 1 lb (at least for Future Meat Technologies, which uses chicken).
This article explains the details of the differences between the 2 types of shotgun seq mentioned.
According to this market report by Grand View Research.
In microbial seq, clinical and research applications, and the NGS market overall, Illumina has a share >65%.
Their paper was published in August of 2022!
2021 paper explaining this tech for “personal use”. Futuristic?
Press release, not including the new center in Brazil.
Fantastic article on this topic by Ginkgo’s very own Sudeep Agarwala and John Han